Sometimes doing science feels like doing magic. Take a fantastical witch brewing eye of toad and nightshade flower in a cauldron, substitute a 1.5 ml tube for the cauldron, AW1 Buffer for the nightshade flower, and blood of junco for the eye of toad, and that’s me.
(And that “eye of ___” thing happens in science too: a few of my herpetologist colleagues have been talking lately about what you can learn from preserved lizard eyes.)
One of the things I do when I capture a junco is to collect a blood sample. I use a sterile needle, collect very little blood, and don’t let the bird go until I’m sure the bleeding has stopped. The birds usually don’t even flinch. They act much more upset when I blow on their chests to look for brood patches (I think it feels cold to them) than they do when I take blood.

Me collecting blood from GRAY. The blood moves up the tiny capillary tube on its own. Photo by M. LaBarbera.
I store the blood in Longmire buffer, which is definitely magic, because it lets you keep blood at room temperature and still use it for lab work months later. Awesome.
In the lab, I extract DNA from the blood. Birds are great for this because they have nucleated red blood cells, and those red blood cells’ nuclei contain DNA, so they have plenty of DNA in their blood. Mammals’ red blood cells don’t contain DNA, so it’s a lot harder to get DNA from mammal blood.
Once I have the DNA I do PCR (Polymerase Chain Reaction) which is a method by which we can create lots and lots of copies of certain pieces of DNA. In my case I make lots of copies of long, repetitive sequences called microsatellites. They might look like: ATCATCATCATCATC…. on and on for 100-300 base pairs.
Microsatellites are useful because all those repetitions occasionally cause the DNA polymerase to slip when it is copying the DNA, so a 20-repetition microsatellite will be copied out as a 19-repetition or a 21-repetition. This happens often enough that you can use microsatellites to distinguish even pretty close relatives: Aunt Susie has a 20-repetition, but Nephew Tommy inherited a 19-repetition. In populations where the animals may all be pretty closely related to each other, this is quite valuable.
A while ago I could have said that microsatellite DNA is noncoding, i.e. it doesn’t “mean” anything; but now it seems like even the most backwater parts of the genome affect something, so I’ll just say that microsats probably don’t mean very much.
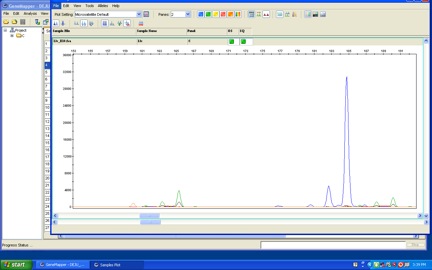
Once I have my PCR products, all those many copies of microsats, I run them through a genotyping machine that reads flashes of fluorescent dyes that I attached to the microsats during PCR. The flashes of dye let the machine figure out how long the microsats are. If all goes well, the output looks something like this:
Here, one of the microsats was dyed green, the other blue. The tallest peaks indicate the length of each microsat (the shorter peaks right before the tallest peaks are “pull-through,” not good data). You can see two different green microsat peaks, indicating two different alleles. (Remember, the birds—and you—have two copies of (almost) all their genes, so they have two copies of each microsat.) In the middle you can see just one big blue peak, indicating that both of this bird’s blue microsat alleles were the same length. This bird was heterozygous for the green microsat and homozygous for the blue microsat.
Once I have these data for lots of microsats and lots of birds, I can compare their amounts of shared and non-shared microsat alleles to see how closely related they are. Fathers should share 50% of their microsats with their chicks, for example.
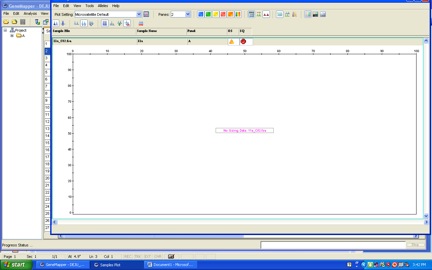
Of course, sometimes the lab work doesn’t go so neatly.
That means I didn’t mix some stuff well enough, and I’ll have to genotype my PCR products again.
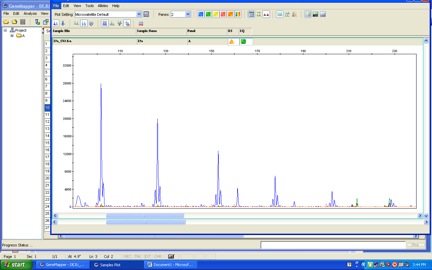
That is just a mess.
I did microsatellite lab work on House Wrens too, and I really like it. Even the birds probably don’t know how related they are to each other, except for parent-offspring and sibling relationships—but soon I will know.


Your posts are always so informative and interesting and I love a good birdy pic. I made myself read through the blood and needles part, just to get to the science explanation. Your title had me worried!
Glad you like it! Thanks for persevering through the blood and needles part :-)
It is really interesting to see more information about the technical side of your lab work.
This non-scientist is confused. You write “Mammals’ red blood cells don’t contain DNA, so it’s a lot harder to get DNA from mammal blood.” I would have thought it would be impossible to get DNA from mammal blood, if the red blood cells of mammals have no DNA.
Great question! I omitted a key bit of information: mammal red blood cells don’t contain DNA, but the white blood cells still do. So there is still some DNA in mammal blood, just less.
Thanks! Makes sense to me now.
I enjoyed your explanation. Have you an interpretation of your data or is it just too soon to ask that? What I really like is your description of when experiments go wrong. It’s so important to recognise this is what can happen but I Know in the lab inconvenient errors might just not be recorded. Understanding the errors is what makes the data valid of course. Beautiful birds, lovely photograph.
Pingback: Featured paper: turkeys help their relatives get lucky | Tough Little Birds
Glad to hear the buffer is working for you! Jon Longmire
Pingback: A new, fuzzier project | Tough Little Birds